Division of Research

Run to R1
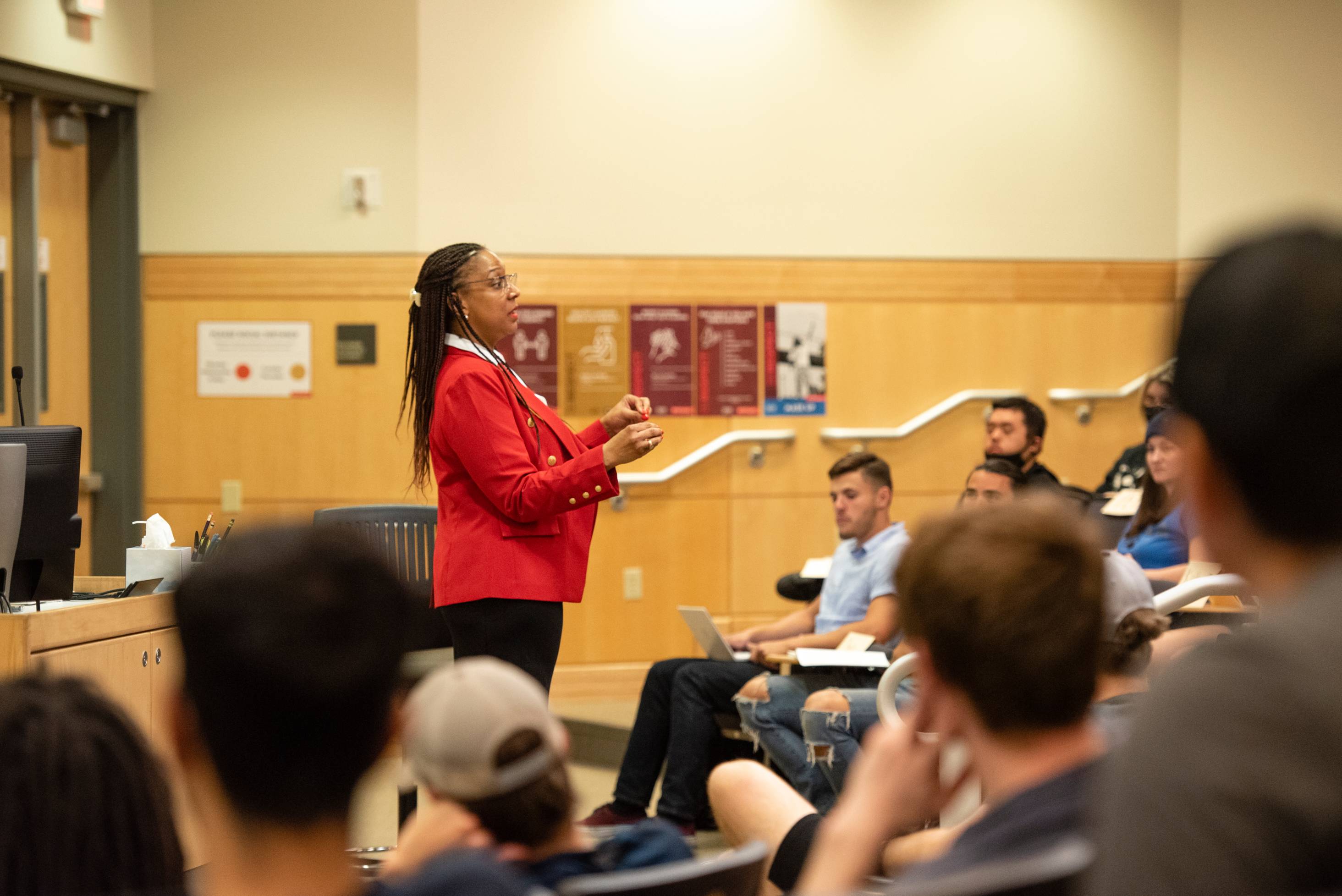
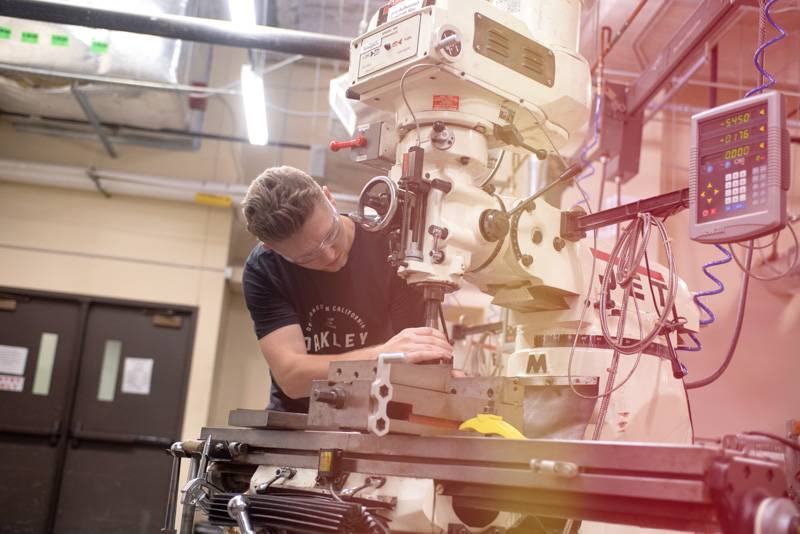
Our faculty and students are creating new knowledge, cultivating tomorrow’s prosperity, and pushing the boundaries of NEXT in every discipline.
$89.7M Total in Awards in FY23
$67M Total Sponsored Programs Expenditure in FY23
Students Resources